Unit Leader

- Harnessing Experimental Animals -
Towards Comprehensive Animal Resource Management, Development, and Technical Support
* Due to the reorganization starting as new centers in April 2018, this laboratory is now belong to the Center for Biosystems Dynamics Research. As for the latest information, please see the following URL below.
> The webpage of Laboratory for Animal Resource Development, Center for Biosystems Dynamics Research

Unit Leader
Hiroshi Kiyonari
2-2-3 Minatojima-minamimachi, Chuo-ku, Kobe, Hyogo 650-0047, Japan
![]()
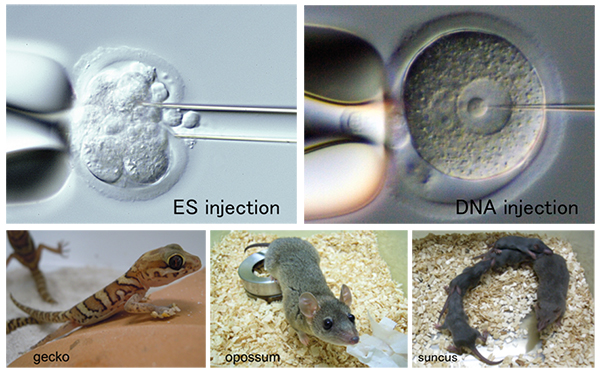
The major mission of the Animal Resource Development Unit is to integrate advanced reproductive and developmental biology technologies and environment for animal research experiments through innovative research in the development of novel experimental animal models and related technical support activities. We utilize our expertise, not only to manage the state-of-the-art SPF rodent facility at RIKEN Kobe Campus, but also to accommodate a variety of animal research needs, including animal material and resource support, technical training, and education for proper conduct of animal research. In recent years, we have established colonies of a Soricomorpha species, suncus (Suncus murinus), a metatherian species, gray short-tail opossum (Monodelphis domestica), and a reptilian species, gecko (Paroedura picta) for prospective distribution services of these new resources to the research community. In addition, in close cooperation with Genetic Engineering Team, we are facilitating joint projects to generate and distribute novel genetically engineered mice with domestic and international biomedical research community, as well as conducting original research projects in live imaging and analyses of early mouse development.

CLST was reorganized into three centers according to the RIKEN 4th Medium-Term Plan from April 1, 2018. For the latest information of Animal Resource Development Unit, please visit the following websites.
> The webpage of Laboratory for Animal Resource Development, Center for Biosystems Dynamics Research [http://www.bdr.riken.jp/en/research/labs/kiyonari-h/index.html]
> If you continue to browse this site, click here.